 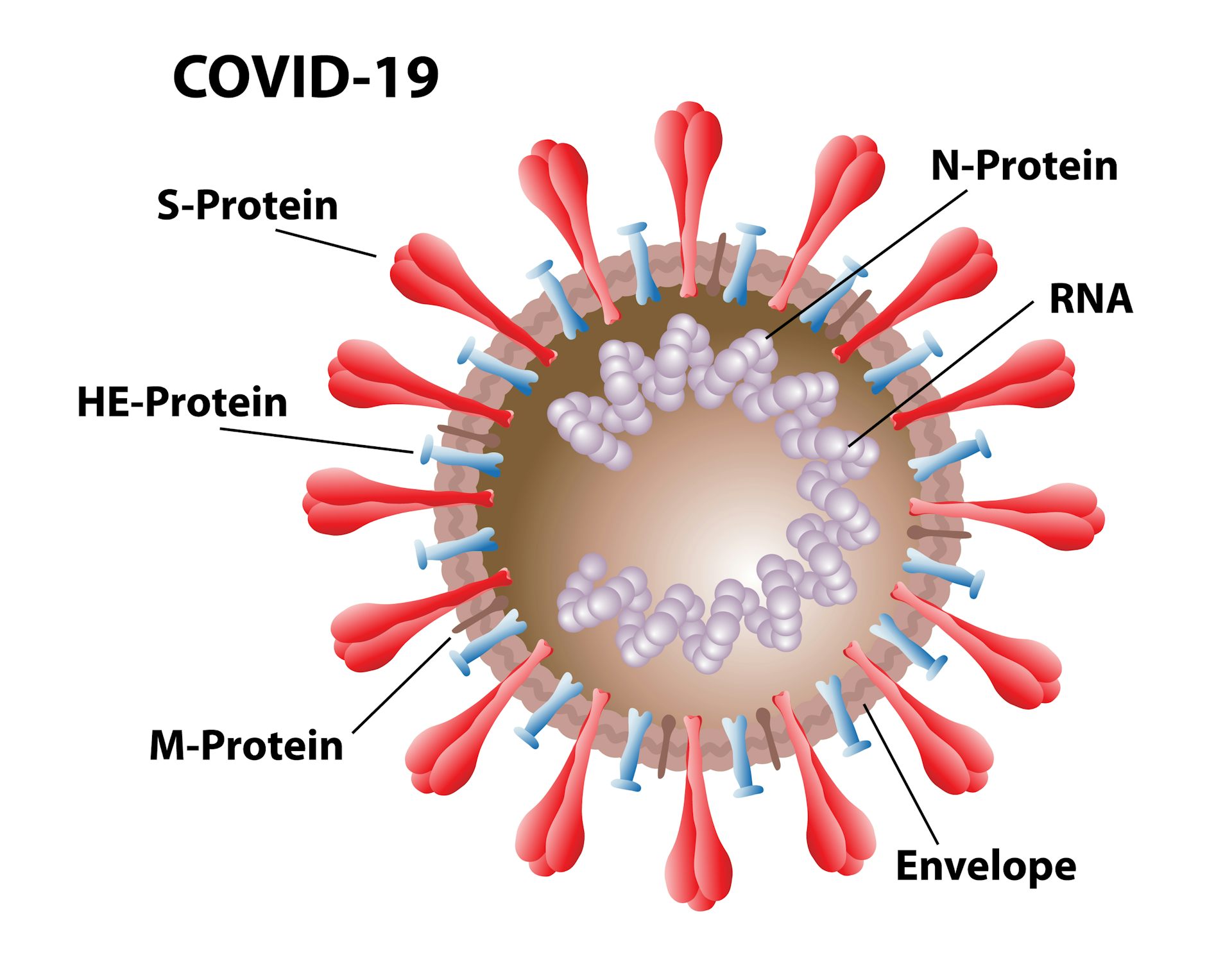
Assessment of its antigenicity and high-resolution EM confirm that this trimer adopts a closed conformation. Combining several of the mutations results in a highly stable S trimer, S-closed, with increased expression that remains stable in the absence of a heterologous trimerization domain that is typically required in soluble S designs 9, 21, 26, 29. In this work, using structure-based design, we find stabilizing mutations in both the S1 and S2 subdomains. Therefore, stabilizing S in its closed (3 RBDs down) state and arresting the first step in the conformational change may result in an improved vaccine immunogen. Although neutralizing antibodies have been mapped to the RBD in the up as well as down state, the antibodies that bind the conserved epitopes on the RBD in the down state were described to have the highest neutralizing potency 4, 6, 13. Comparison of the structure of SARS-CoV-2 S-2P 9, 26 with that of native SARS-CoV-2 S 27, 28 shows that the former adopts a more open conformation with one or more of the RBDs in the ‘up’ conformation. However, the SARS-CoV-2 S protein carrying these substitutions and additional furin site mutations (S-2P) remains unstable and strategies to further improve its stability have recently been described 12, 23, 24, 25. Stabilization of the hinge loop of the S proteins of SARS-CoV and MERS-CoV has been achieved by mutation of two consecutive residues in the S2 subunit between the central helix (CH) and heptad repeat 1 (HR1) 21, 22 to proline and this approach (2P) has successfully been applied to the SARS-CoV-2 S protein 9. A particularly successful approach to enhance prefusion stability was shown to be the stabilization of the so-called hinge loop preceding the central helix (CH), which has been applied to a range of viral fusion glycoproteins 15, 16, 17, 18, 19, 20. In recent years, efforts have been made to stabilize various class I fusion proteins through structure-based design (for a review see ref. Prefusion stabilization typically increases the recombinant expression of viral glycoproteins, which facilitates the production of protein for (subunit) vaccines and improves the immune response elicited by recombinant protein and viral vector vaccines 14. The prefusion conformation of S as present on the infectious particle contains the epitopes for neutralizing antibodies and thus holds most promise as a vaccine immunogen 3, 4, 5, 6, 12, 13. Like other class I fusion proteins, the SARS-CoV-2 S protein is intrinsically metastable as a consequence of its ability to undergo extensive conformational changes that are required to drive fusion. The receptor-binding site, which can bind to human angiotensin-converting enzyme 2 (ACE2), is transiently exposed in the ‘up’ conformation. 
In the prefusion state, the S-protein’s RBD domains alternate between open (‘up’) and closed (‘down’) conformations. For infection, S requires proteolytic cleavage by a furin-like protease between the S1 and S2 subunits (S1/S2), and by TMPRSS2 at a conserved site directly preceding the fusion peptide (S2’) 10, 11. The S1 subunit comprises two distinct domains: an N-terminal domain (NTD) and a host cell receptor-binding domain (RBD). S is a large, trimeric glycoprotein that mediates both binding to host cell receptors and fusion of virus and host cell membranes, through its S1 and S2 subunits, respectively 7, 8, 9. The viral surface spike (S) protein is a key target for prophylactic measures as it is critical in the viral life cycle and the primary target of neutralizing antibodies 3, 4, 5, 6. Highly stable and well producing S protein and the increased understanding of S protein structure will support vaccine development and serological diagnostics.ĭevelopment of effective preventative interventions against the SARS-CoV-2 virus that causes the ongoing COVID-19 pandemic 1, 2 is urgently needed. The cryo-EM structure reveals a correctly folded, predominantly closed pre-fusion conformation. We combine a minimal selection of mostly interprotomeric mutations to create a stable S-closed variant with a 6.4-fold higher expression than the parental construct while no longer containing a heterologous trimerization domain. We identify regions critical for stability: the heptad repeat region 1, the SD1 domain and position 614 in SD2. To support ongoing vaccine development efforts, we report the structure-based design of soluble S trimers with increased yields and stabilities, based on introduction of single point mutations and disulfide-bridges. Due to intrinsic instability typical of class I fusion proteins, S tends to prematurely refold to the post-fusion conformation, compromising immunogenic properties and prefusion trimer yields. The trimeric spike (S) protein of SARS-CoV-2 is the primary focus of most vaccine design and development efforts. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
